-Search query
-Search result
Showing 1 - 50 of 153 items for (author: kubo & s)

EMDB-36223: 
Cryo-EM structure of the GI.4 Chiba VLP complexed with the CV-1A1 Fv-clasp
Method: single particle / : Hosaka T, Katsura K, Kimura-Someya T, Someya Y, Shirouzu M

EMDB-17171: 
CTE typeI tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17173: 
CTE typeIII tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17174: 
CTE typeII tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17175: 
TMEM106B Fold1-s filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17176: 
TMEM106B Fold I-d filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17177: 
Ab typeII filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17178: 
CTE typeI tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17179: 
TypeII tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17180: 
CTE typeIII tau filament
Method: helical / : Tetter S, Qi C, Ryskeldi-Falcon B, Scheres SHW, Goedert M

EMDB-17181: 
PHF tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8ot6: 
CTE typeI tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

PDB-8ot9: 
CTE typeIII tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8otc: 
CTE typeII tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8otd: 
TMEM106B Fold1-s filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

PDB-8ote: 
TMEM106B Fold I-d filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

PDB-8otf: 
Ab typeII filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8otg: 
CTE typeI tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
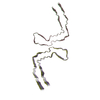
PDB-8oth: 
TypeII tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
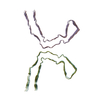
PDB-8oti: 
CTE typeIII tau filament
Method: helical / : Tetter S, Qi C, Ryskeldi-Falcon B, Scheres SHW, Goedert M
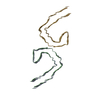
PDB-8otj: 
PHF tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-41171: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, wild-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L

EMDB-41172: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, variant-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L
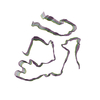
PDB-8tdn: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, wild-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L
PDB-8tdo: 
Cryo-EM structure of cardiac amyloid fibril from a variant ATTR I84S amyloidosis patient-3, variant-type morphology
Method: helical / : Nguyen BA, Singh V, Saelices L

EMDB-35500: 
Cryo-EM structure of the gastric proton pump with bound DQ-02
Method: single particle / : Abe K, Yokoshima S, Yoshimori A

EMDB-35501: 
Cryo-EM structure of the gastric proton pump with bound DQ-06
Method: single particle / : Abe K, Yokoshima S, Yoshimori A

EMDB-35502: 
Cryo-EM structure of the gastric proton pump with bound DQ-18
Method: single particle / : Abe K, Yokoshima S, Yoshimori A

EMDB-36424: 
Cryo-EM structure of the gastric proton pump with bound DQ-21
Method: single particle / : Abe K, Yokoshima S, Yoshimori A

PDB-8ijv: 
Cryo-EM structure of the gastric proton pump with bound DQ-02
Method: single particle / : Abe K, Yokoshima S, Yoshimori A

PDB-8ijw: 
Cryo-EM structure of the gastric proton pump with bound DQ-06
Method: single particle / : Abe K, Yokoshima S, Yoshimori A

PDB-8ijx: 
Cryo-EM structure of the gastric proton pump with bound DQ-18
Method: single particle / : Abe K, Yokoshima S, Yoshimori A

PDB-8jmn: 
Cryo-EM structure of the gastric proton pump with bound DQ-21
Method: single particle / : Abe K, Yokoshima S, Yoshimori A

EMDB-33643: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab816
Method: single particle / : Uchikubo T, Shirouzu M

EMDB-33644: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab803
Method: single particle / : Uchikubo T, Shirouzu M

PDB-7y6l: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab816
Method: single particle / : Uchikubo T, Shirouzu M

PDB-7y6n: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab803
Method: single particle / : Uchikubo T, Shirouzu M

EMDB-29666: 
96-nm repeat unit of intact doublet microtubule from Tetrahymena thermophila CFAP77AB-KO mutant
Method: subtomogram averaging / : Black CS, Legal T, Bui KH

EMDB-29667: 
96-nm repeat unit of intact doublet microtubule from Tetrahymena thermophila strain CU428
Method: subtomogram averaging / : Black CS, Legal T, Bui KH

EMDB-29685: 
48-nm doublet microtubule from Tetrahymena thermophila strain CU428
Method: single particle / : Black CS, Kubo S, Yang SK, Bui KH

EMDB-29692: 
48-nm doublet microtubule from Tetrahymena thermophila strain K40R
Method: single particle / : Black CS, Kubo S, Yang SK, Bui KH

EMDB-29693: 
96-nm doublet microtubule from combined Tetrahymena thermophila strains CU428 and K40R
Method: single particle / : Black CS, Kubo S, Yang SK, Bui KH

PDB-8g2z: 
48-nm doublet microtubule from Tetrahymena thermophila strain CU428
Method: single particle / : Black CS, Kubo S, Yang SK, Bui KH

PDB-8g3d: 
48-nm doublet microtubule from Tetrahymena thermophila strain K40R
Method: single particle / : Black CS, Kubo S, Yang SK, Bui KH

EMDB-34588: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 1 (3.2 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34589: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 2 (3.9 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34590: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 4 (3.9 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Sato S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34591: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 1 (4.7 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34592: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 2 (4.8 angstrom resolution)
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Shin S, Uchikubo-Kamo T, Shirouzu M, Umehara T

EMDB-34593: 
Cryo-EM structure of the p300 catalytic core bound to the H4K12acK16ac nucleosome, class 3
Method: single particle / : Kikuchi M, Morita S, Wakamori M, Sato S, Uchikubo-Kamo T, Shirouzu M, Umehara T
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
